Definition 8.5. [BLL98, p. 8] The species E of sets (french: ensemble) is
defined by E[U] = 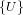 for any finite set U. For any bijection σ : U → V , E[σ]
is given by mapping the only element U ∈ E[U] to the only element V ∈ E[V ].
for any finite set U. For any bijection σ : U → V , E[σ]
is given by mapping the only element U ∈ E[U] to the only element V ∈ E[V ].
Definition 8.5. [BLL98, p. 8] The species E of sets (french: ensemble) is
defined by E[U] = 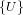 for any finite set U. For any bijection σ : U → V , E[σ]
is given by mapping the only element U ∈ E[U] to the only element V ∈ E[V ].
for any finite set U. For any bijection σ : U → V , E[σ]
is given by mapping the only element U ∈ E[U] to the only element V ∈ E[V ].
Originally, we wanted to extend the existing Set constructor from the Aldor library. Unfortunately, Set does not guarantee that in
a: List L := [1,2,3];
s: Set L := [l for l in a];
b: List L := [l for l in s];
From a mathematical point of view there is no problem at all. However, computationally, it might be desirable to be able to tell something about the order of the set elements, in order to go conveniently back and forth between Set and List. In fact, for SetSpecies we would like to be able to turn a list into a set in constant time, since in some cases we know that the list does not contain duplicate elements.
Since, we cannot access the internal data structure of Set, we cannot simply extend Set and thus must provide SetSpecies.
Since we currently do not need much of the functionality of Set and, in fact, SetSpecies should in many respects behave like the List data structure (not the combinatorial species), we provide only the functionality that is needed.
First of all, the following code should give a = b.
a: List L := [1,2,3];
s: SetSpecies L := set(a);
b: List L := s :: List(L);
Furthermore, for any F of type
(L: LabelType) -> CombinatorialSpecies L
(s :: List L) = (t :: List L)
Such a deterministic behavior is vital for reliable ranking and unranking algorithms.
The argument against using List is that as combinatorial species List and SetSpecies are quite different. And, additionally, by using List we would lose the information that the list in question has no duplicate elements.
Note that SetSpecies and CombinatorialSpecies should be defined in the same file since they mutually depend on each other.
Usage
s: SetSpecies Integer := [3, 5, 2];
for x in structures(s) repeat stdout << x << newline;
Description
The Set constructor seen as a combinatorial species.
bracket: Tuple L -> %
bracket(t: Tuple L): % == {
l: List L == [t];
s: % := empty;
while not empty? l repeat {
s := union(first l, s);
l := rest l;
}
per reverse! rep s;
}
Exports of SetSpecies
Remember that # must not appear in the first column, otherwise it might be misinterpreted by the compiler as a system command.
The following might lead to non-sets if applied to bad data.
Export of SetSpecies
cons: (L, %) -> %
Description
Adds a new element to the set.
The function inserts a new element in such a way that the new element will have index 1.
Remarks
The function does not check whether the element is already in the set and thus is potentially type-unsafe.
Export of SetSpecies
set: List L -> %
Description
Turns a list into a set species.
The function turns a list into a set without checking whether all elements in the list are different.
Remarks
The function does not check whether given list has pairwise different elements and thus is potentially type-unsafe.
The generating series is ex.
And the isomorphism type generating series is 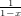 .
.
The cycle index series ZE of the species E of sets is given by
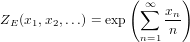 | (12) |